PET-BIDS on SPICE
Raw data from BMIC
PET produce different kind of data uploaded from BMIC in folders labeled:
- PT (Positon Tomography)
- CT (Computed Tomography)
- Blood
- PT_fr (frame rate motion corrected PT)
- PT_er (event rate motion corrected PT)
You will find these folders in a sub-folder created for the radioligand used. Actual filenames of the raw DICOMs will depend on your protocol name, but essentially the raw data in your project on SPICE will be structured like this:
/data/projects/yourproject/raw/pet/sub-XXX/ses-XX/ucb-j
├── PT
| ├ 0000001.dcm
| ├ 0000002.dcm
| ├ ....
| └─0022500.dcm
└── CT
├ 0000001.dcm
├ 0000002.dcm
├ ....
└─0022500.dcm
Based on the availability of the raw data folder /data/projects/yourproject/raw/bmic, a PET tab will show up that includes all tools necessary to BIDSify PET data. If your project folder also contains a raw MRI folder /data/projects/yourproject/raw/mri, you will also have an MRI tab with similar functionalities. See the MRI page for how to convert your MRI data to BIDS.
See general steps (0 to 2) on: BIDS on SPICE
There are some common steps you should go through for making any imaging data to BIDS, see link above.
3. Analyze your data
Open the URL in your local browser. If this is your first time, you will see the window below. You are prompted to select one representative session from your data. This session will be analyzed with the dcm2bids_helper function. Make sure to use a session that is complete and contains all sequences you want in your final data set.
This function goes through the DICOM files in the selected session and export their unique DICOM meta data to a JSON file for each unique sequence. These are saved in the cir-utils folder on SPICE under dcm2bids_helper/helper. The config is saved in the cir-utils/tabs/pet-bids folder as dcm2bids_config_pet.json
Note that while you can use dcm2niix to convert your CT sequence to BIDS, there is no official BIDS for CT yet. File naming from CT conversion will follow the BIDS Extension Proposal for CT BEP-024. If you need your BIDS dataset to be fully BIDS compliant as checked by the BIDS validator, do not convert your CT data to BIDS.

Clicking Analyze DICOM fields will start the analysis. Depending on how many and what sequences are in the selected session, this will take a couple of minutes to complete. The ouput from dcm2bids_helper is printed in the faux terminal window on the page. When the analysis is done, a summary of the most commonly used fields are printed in a table in the browser so that you can use these to build your config file for dcm2bids. These are:
Series number: The order the sequence was run in your protocol.Series description: The name of the sequence in your protocol.Protocol name: The name of the protocol, often including the name of your project and tracerModality: PT for PET and CT for CTImage type: A list of keywords describing the sequence, e.g.ORIGINAL,PRIMARY,PERFUSIONRadiopharmaceutical: The name of the radiopharmaceutical in your dicom header, note that if this is incorrect, you should edit your json files to included updated tracer info
You can select and analyze different sessions for defining your config file, but easiest for you is to select a session that you know contain all the sequences you are interested in. Check the box Force dcm2bids_helper to overwrite the current JSON helper files with new ones.
4. Build config file
The input
When dcm2bids_helper is done, the section 2. Specify config shows up. This section summarize the DICOM fields from the JSON files created by dcm2bids_helper (i.e. your representative session). Now, you can use what is in your data to construct a config file for dcm2bids so it knows what inputs to look for in your raw data and what outputs to genarate as BIDS. This is an important step that can require some detective work, especially if you skipped step 0 above, but you only need to do this once.
If you ran the helper before, this table will show the summary from the last session you analyzed. In the image below is an example of a session that has one CT sequence and two PET sequences, the original dynamic data and the data with an additional motion correction.
The left columns under INPUT are used to define the inputs for dcm2bids. That is, what do you want dcm2bids to look for in your data. Click on the fields in the table to select them to be included as a criteria for that sequence in the config file. You must select at least one field as a criteria in each sequence you want included.
If you want dcm2bids to skip a sequence, simply do not select any criteria for that sequence and it will be ignored, and not included in the BIDS output.

Note that the two PET sequences and the CT sequence have the same Protocol name, so make sure you select a unique identifier (in this case Series description) or unique combination of identifiers for every included sequence.
The output
For the output section you MUST specify the datatype and suffix as indicated by an angry asterisk. For data from BMIC, datatype is either pet, or anat. The suffix MUST match the datatype, and selecting the datatype will limit the options for the suffic. For projects with only one tracer, you do not have to fill in trc. However, in projects with multiple tracers, failing to fill in trc will lead to issues, as both sequences from the same subject and session will be saved under the same name. Similarly, different reconstructions should use distinct rec flags
In the image below, you can see we have specified datatype pet and suffix pet for the PET sequence with trc as ucbj, and datatype ct with suffix ct for the CT sequence. This will result in the PET sequence being saved as sub-XXX_ses-XX_trc-ucbj_pet.nii.gz and the motion correct PET being saved as sub-XXX_ses-XX_trc-ucbj_rec-acdyn_pet.nii.gz. The CT sequence will be saved as sub-XXX_ses-XX_ct.nii.gz in the BIDS output, but keep in mind that there is no official CT BIDS yet.

The other output fields are optional but can be used to further specify unique output file names. For example, many fMRI protocols include a task that should then be specified. Consult the BIDS documentation for the use of different entities, in short:
- Task name
task: The name of the task performed during the scan, e.g.restfor resting state PET - Run
run: The run number of the sequence. Useful if you have more than one run of the same sequence in your protocol
When you have specified a unique input as criteria, and a unique output for the resulting BIDS, for each sequence you want to include you click Generate config file to write your specifications to dcm2bids_config_pet.json in the cir-utils folder on SPICE. This is the config file that will be used as input for the BIDS conversion, which uses a procedure similar to dcm2bids. This file will be loaded automatically next time you open this page, but you can also load it manually by clicking Load config file.
If you do not have a complete, representative session with all the sequences you want to include in your BIDS output, you can still use Append to config to append a new sequence input criteria and output structure to the config file while keeping the existing specifications.
You can also edit the config file manually if you want, in the tab Config Editor - but be careful to keep the JSON format.
5. Inspect your config file
You can view and make manual edits to your config file in the tab Config Editor. Be careful to keep the JSON format if you make manual edits here. It may be a good idea to keep a backup of your config file, both for book keeping and especially before making manual edits.
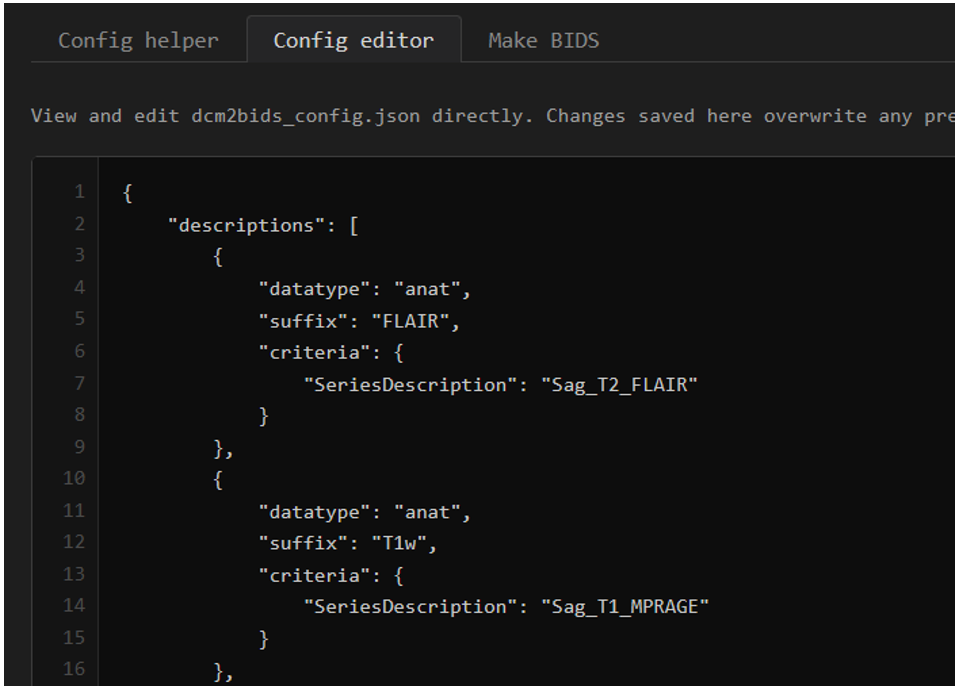
What the bare bones of the config file look like - a JSON file with a list of dictionaries. Here an example from MRI is shown.
6. Check subjects and sessions
On the final tab Make BIDS you first check that paths you are using in your project folder are correct and then click Discover sessions to list all the sessions available for you to BIDSify.

A table will show all subjects and sessions available in your raw data folder. You can specify which subjects and sessions to include in the BIDS output by checking the boxes in the table. Generally CIR will deliver your data in a pseduo-BIDS format but you can rename a subject and session if your raw data is not matching what you need for your BIDS output. Your recode-key will be saved as session_recode_pet.csv in the cir-utils/tabs/pet-bids folder on SPICE for you to keep track of how you have renamed your data. Below is an example of MRI data with a participant ID that needs recoding.
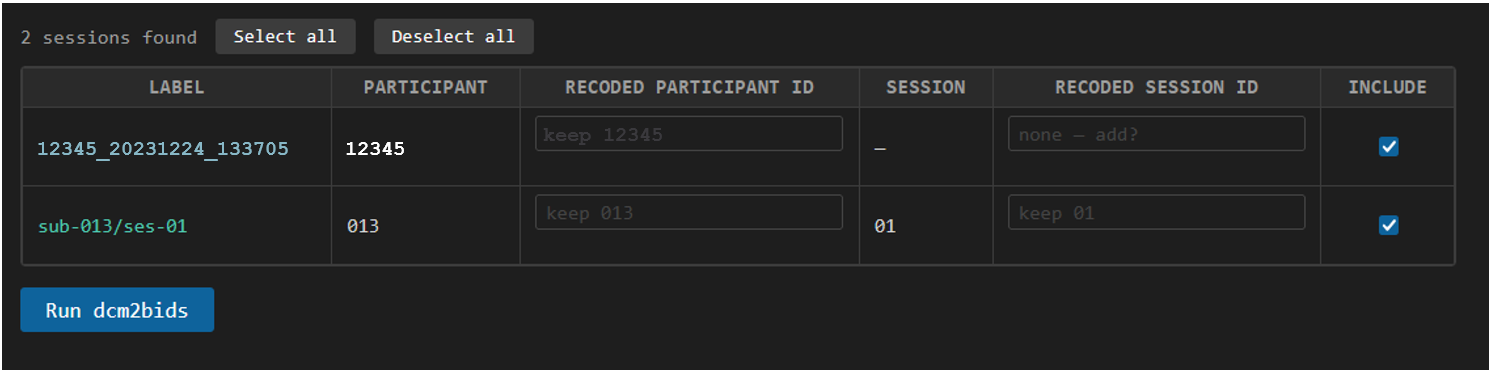
In the image above we see two sessions discovered, one subject is saved as an ID and date-time 12345_20231224_133705, a legacy format for MRI data at MRC. The participant ID is correctly extracted, but doesn't match our other subject and there is no session number.
We simply type in a new subject ID and session number in the table to recode this session to match the BIDS output we want. In this case, we recode the subject to sub-012 and session to ses-01. This is saved in the recode key so that next time we open this page, the same session is recoded to the same subject and session ID. Three digit subject IDs and two digit session numbers, both with leading zero-padding, are recommended.
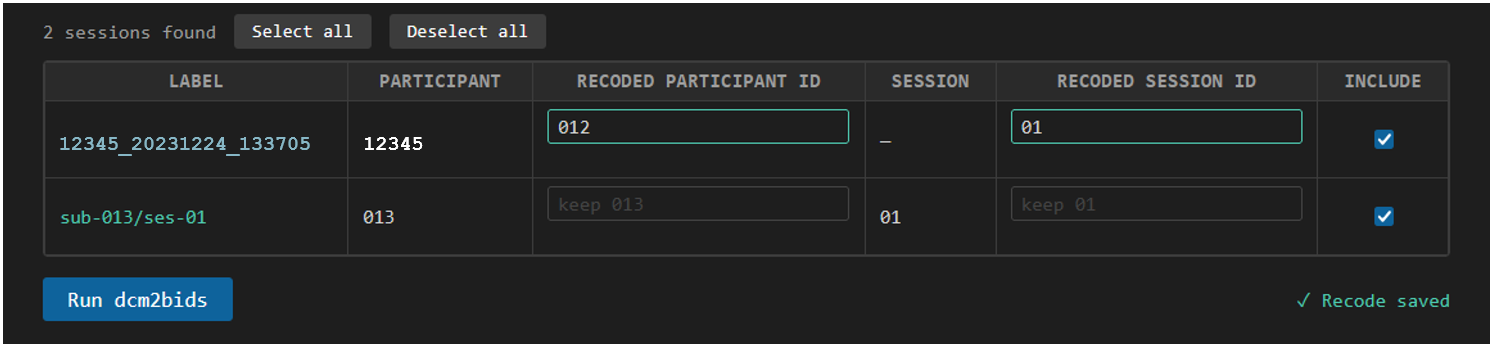
7. Run PET BIDS
Finally, you click Run PET BIDS to start the processing. This will run pypet2bids for any PET sequences and dcm2niix for any CT sequences. The conversion is executed using both the config and the recoding csv.
Depending on how many sessions you have selected and how many sequences are in your protocol, this can take a while to complete. The output from the conversion is printed in the faux terminal window on the page so you can follow the progress and check for any errors.
This process is designed to be able to run regularly during your project. No need to wait until you have collected all your data to start organizing it as BIDS. Rather, start early to identify and correct any issues with your config file or mismatch naming in your protocol or raw data.
Your config file and recode key are saved in the cir-utils folder in your project folder on SPICE (config in cir-utils/dcm2bids_config_pet.json and recode key in cir-utils/tabs/pet-bids/sessions_recode_pet.json) and running the BIDS conversion does not overwrite already existing BIDS files unless you tell it to (by checking --clobber).
8. Celebrate
Congratulations!! 🎉 You have now organized your data as BIDS and can start using common pre-processing pipelines and tools for your data analysis. You are contributing to making your data more FAIR and reusable for other researchers, and you are saving time for yourself and others in the future by not having to re-invent the wheel for organizing your data for every new project. You can also share your config file with other researchers at CIR to further promote efficient data handling and sharing of best practices between projects.